An elegant way to count number of negative elements in a vector?
Solution 1
You want to read 'An Introduction to R'. Your answer here is simply
sum( x < 0 )
which works thanks to vectorisation. The x < 0 expression returns a vector of booleans over which sum() can operate (by converting the booleans to standard 0/1 values).
Solution 2
There is a good answer to this question from Steve Lianoglou How to identify the rows in my dataframe with a negative value in any column?
Let me just replicate his code with one small addition (4th point).
-
Imagine you had a data.frame like this:
df <- data.frame(a = 1:10, b = c(1:3,-4, 5:10), c = c(-1, 2:10)) -
This will return you a boolean vector of which rows have negative values:
has.neg <- apply(df, 1, function(row) any(row < 0)) -
Here are the indexes for negative numbers:
which(has.neg) -
Here is a count of elements with negative numbers:
length(which(has.neg))
Related videos on Youtube
Carey
Updated on November 01, 2020Comments
-
Carey over 3 years
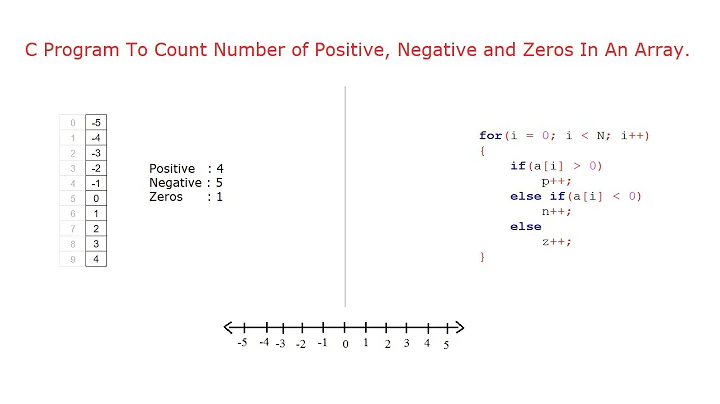
I have a data vector with 1024 values and need to count the number of negative entries. Is there an elegant way to do this without looping and checking if an element is <0 and incrementing a counter?
-
Matthew Lundberg over 11 yearsThe vector is not converted to numeric (for later versions of R anyway) - sum works on logical directly. This can be easily observed.
x <- rep(c(T,F), 2^30-1)causes my R session RSS to increase to ~8G.sum(x)does not increase this. However,sum(as.numeric(x))causes the RSS to increase to ~25G (and of course returns the same value). -
 Jeffrey Evans over 6 yearsThe OP has a vector so, something like length(x[x < 0]) would work.
Jeffrey Evans over 6 yearsThe OP has a vector so, something like length(x[x < 0]) would work. -
 Andrii over 6 yearsIf data set has NA values the command should be sum(x < 0, na.rm = TRUE). Otherwise it returns NA as a result
Andrii over 6 yearsIf data set has NA values the command should be sum(x < 0, na.rm = TRUE). Otherwise it returns NA as a result