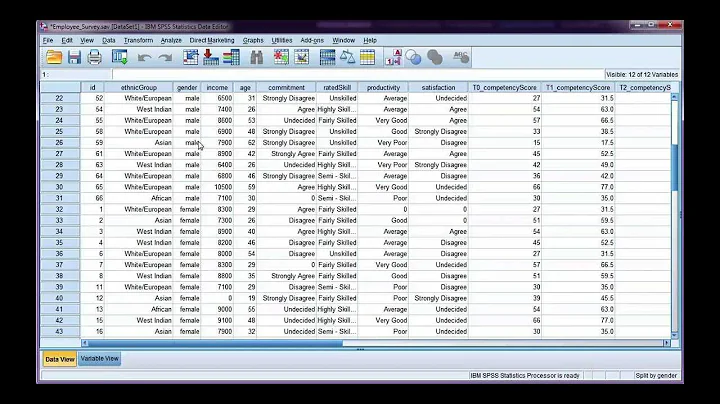
How do I break up a file like split to stdout for piping to a command?
Solution 1
I ended up with something that's seemingly gross, if there's a better way please post it:
#!/bin/sh
DONE=false
until $DONE; do
for i in $(seq 1 $2); do
read line || DONE=true;
[ -z "$line" ] && continue;
lines+=$line$'\n';
done
sql=${lines::${#lines}-10}
(cat "Header.sql"; echo "$sql";) | sqlcmd
#echo "--- PROCESSED ---";
lines=;
done < $1
Run with ./insert.sh "File.sql" 100 where the 100 is the number of lines to process at a time.
Solution 2
I think the easiest way to do this is:
while IFS= read -r line; do
{ printf '%s\n' "$line"; head -n 99; } |
other_commands
done <database_file
You need to use read for the first line in each section as there appears to be no other way to stop when the end of the file is reached. For more information see:
- Check if pipe is empty and run a command on the data if it isn't
- How to pipe output from one process to another but only execute if the first has output?
Solution 3
Basically, I'm looking for
splitthat will output tostdout, not files.
If you have access to gnu split, the --filter option does exactly that:
‘--filter=command’
With this option, rather than simply writing to each output file, write
through a pipe to the specified shell command for each output file.
So in your case, you could either use those commands with --filter, e.g.
split -l 100 --filter='{ cat Header.sql; cat; } | sqlcmd; printf %s\\n DONE' infile
or write a script, e.g. myscript:
#!/bin/sh
{ cat Header.sql; cat; } | sqlcmd
printf %s\\n '--- PROCESSED ---'
and then simply run
split -l 100 --filter=./myscript infile
Solution 4
GNU Parallel is made for this:
cat bigfile | parallel --pipe -N100 yourscript
It will default to running 1 job per CPU core. You can force running a single job with '-j1'.
Version 20140422 includes a fast version that can deliver 3.5 GB/s. The price is that it cannot deliver the exact 100 lines, but if you know the approximate line length you can set --block to 100 times that (here I assume the line length is close to 500 bytes):
parallel --pipepart --block 50k yourscript :::: bigfile
Solution 5
_linc() ( ${sh-da}sh ${dbg+-vx} 4<&0 <&3 ) 3<<-ARGS 3<<\CMD
set -- $( [ $((i=${1%%*[!0-9]*}-1)) -gt 1 ] && {
shift && echo "\${inc=$i}" ; }
unset cmd ; [ $# -gt 0 ] || cmd='echo incr "#$((i=i+1))" ; cat'
printf '%s ' 'me=$$ ;' \
'_cmd() {' '${dbg+set -vx ;}' "$@" "$cmd" '
}' )
ARGS
s= ; sed -f - <<-INC /dev/fd/4 | . /dev/stdin
i_cmd <<"${s:=${me}SPLIT${me}}"
${inc:+$(printf '$!n\n%.0b' `seq $inc`)}
a$s
INC
CMD
The above function uses sed to apply its argument list as a command string to an arbitrary line increment. The commands you specify on the command line are sourced into a temporary shell function which is fed a here document on stdin consisting of every increment's step worth of lines.
You use it like this:
time printf 'this is line #%d\n' `seq 1000` |
_linc 193 sed -e \$= -e r \- \| tail -n2
#output
193
this is line #193
193
this is line #386
193
this is line #579
193
this is line #772
193
this is line #965
35
this is line #1000
printf 'this is line #%d\n' `seq 1000` 0.00s user 0.00s system 0% cpu 0.004 total
The mechanism here is very simple:
i_cmd <<"${s:=${me}SPLIT${me}}"
${inc:+$(printf '$!n\n%.0b' `seq $inc`)}
a$s
That's the sed script. Basically we just printf $increment * n;. So if you set your increment to 100 printf will write you a sed script consisting of 100 lines that say only $!n, one insert line for the top end of the here-doc, and one append for the bottom line - that's it. Most of the rest just handles options.
The next command tells sed to print the current line, delete it, and pull in the next one. The $! specifies that it should only try on any line but the last.
Provided only an incrementer it will:
printf 'this is line #%d\n' `seq 10` | ⏎
_linc 3
#output
incr #1
this is line #1
this is line #2
this is line #3
incr #2
this is line #4
this is line #5
this is line #6
incr #3
this is line #7
this is line #8
this is line #9
incr #4
this is line #10
So what's happening behind the scenes here is the function is set to echo a counter and cat its input if not provided a command string. If you saw it on the command line it would look like:
{ echo "incr #$((i=i+1))" ; cat ; } <<HEREDOC
this is line #7
this is line #8
this is line #9
HEREDOC
It executes one of these for every increment. Look:
printf 'this is line #%d\n' `seq 10` |
dbg= _linc 3
#output
set -- ${inc=2}
+ set -- 2
me=$$ ; _cmd() { ${dbg+set -vx ;} echo incr "#$((i=i+1))" ; cat
}
+ me=19396
s= ; sed -f - <<-INC /dev/fd/4 | . /dev/stdin
i_cmd <<"${s:=${me}SPLIT${me}}"
${inc:+$(printf '$!n\n%.0b' `seq $inc`)}
a$s
INC
+ s=
+ . /dev/stdin
+ seq 2
+ printf $!n\n%.0b 1 2
+ sed -f - /dev/fd/4
_cmd <<"19396SPLIT19396"
this is line #1
this is line #2
this is line #3
19396SPLIT19396
+ _cmd
+ set -vx ; echo incr #1
+ cat
this is line #1
this is line #2
this is line #3
_cmd <<"19396SPLIT19396"
REALLY FAST
time yes | sed = | sed -n 'p;n' |
_linc 4000 'printf "current line and char count\n"
sed "1w /dev/fd/2" | wc -c
[ $((i=i+1)) -ge 5000 ] && kill "$me" || echo "$i"'
#OUTPUT
current line and char count
19992001
36000
4999
current line and char count
19996001
36000
current line and char count
[2] 17113 terminated yes |
17114 terminated sed = |
17115 terminated sed -n 'p;n'
yes 0.86s user 0.06s system 5% cpu 16.994 total
sed = 9.06s user 0.30s system 55% cpu 16.993 total
sed -n 'p;n' 7.68s user 0.38s system 47% cpu 16.992 total
Above I tell it to increment on every 4000 lines. 17s later and I've processed 20 million lines. Of course the logic isn't serious there - we only read each line twice and count all of their characters, but the possibilities are pretty open. Also if you look closely you might notice it's seemingly the filters providing the input that are taking the majority of the time anyway.
Related videos on Youtube
Ehryk
I'm a Computer (Web) Programmer/Analyst based in Anchorage, AK and Minneapolis, MN. I use (among other things) ASP.NET, C# and SQL Server. I build things. Bicycles, computers, websites, guitars, cars, motorcycles, sound sytems... lots of things. Resume: http://ericmenze.com Personal Website: http://ehryk.com Pause your videos at specific locations: http://pauseforlater.com Calculate and build spoked bicycle wheels: http://wheelspoking.com See activity specific analysis of your GPX Files: http://gpxdataanalyzer.com Tool to open command/powershell prompts from any location (Windows): https://github.com/Ehryk/ContextMenuTools
Updated on September 18, 2022Comments
-
Ehryk over 1 year
I have a large
.sqlfile full ofSELECTstatements that contain data I want to insert into my SQL Server database. I'm looking for how I could basically take the file's contents, 100 lines at a time, and pass it to the commands I have set to do the rest.Basically, I'm looking for
splitthat will output tostdout, not files.I'm also using CygWin on Windows, so I don't have access to the full suite of tools.
-
 Admin about 10 yearsHave you looked at using
Admin about 10 yearsHave you looked at usingBULK INSERT? Separate the data from the SQL statement.
-
-
Graeme about 10 yearsI'm not sure exactly what assumptions are safe with SQL, but for general safety you should do
IFS= read -r line. Consider the different betweenecho ' \t\e\s\t ' | { read line; echo "[$line]"; }andecho ' \t\e\s\t ' | { IFS= read -r line; echo "[$line]"; }. Alsoechois not safe with arbitrary strings (egline="-n"; echo "$line"), it is safer to useprintf '%s\n. -
keen over 7 yearsit's worth noting that the shear complexity of the shell magic in this makes it not portable - it certainly doesnt run on bash4 on osx 10.9. :) it wants to expand to use
dash, andsed -f -doesnt make bsd sed happy either... not to mention having to pull the heredoc markers back to ^...